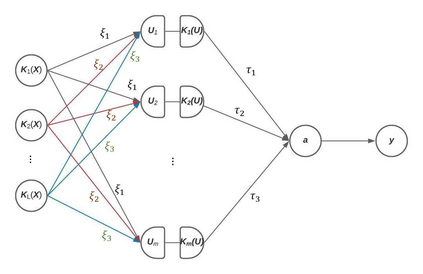
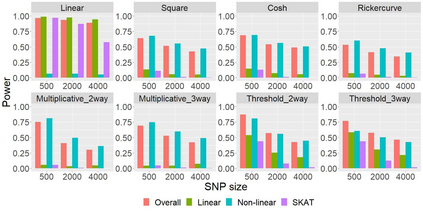
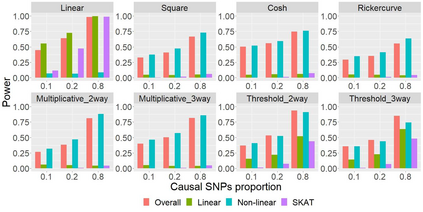
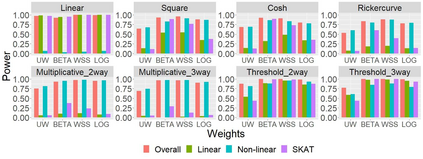
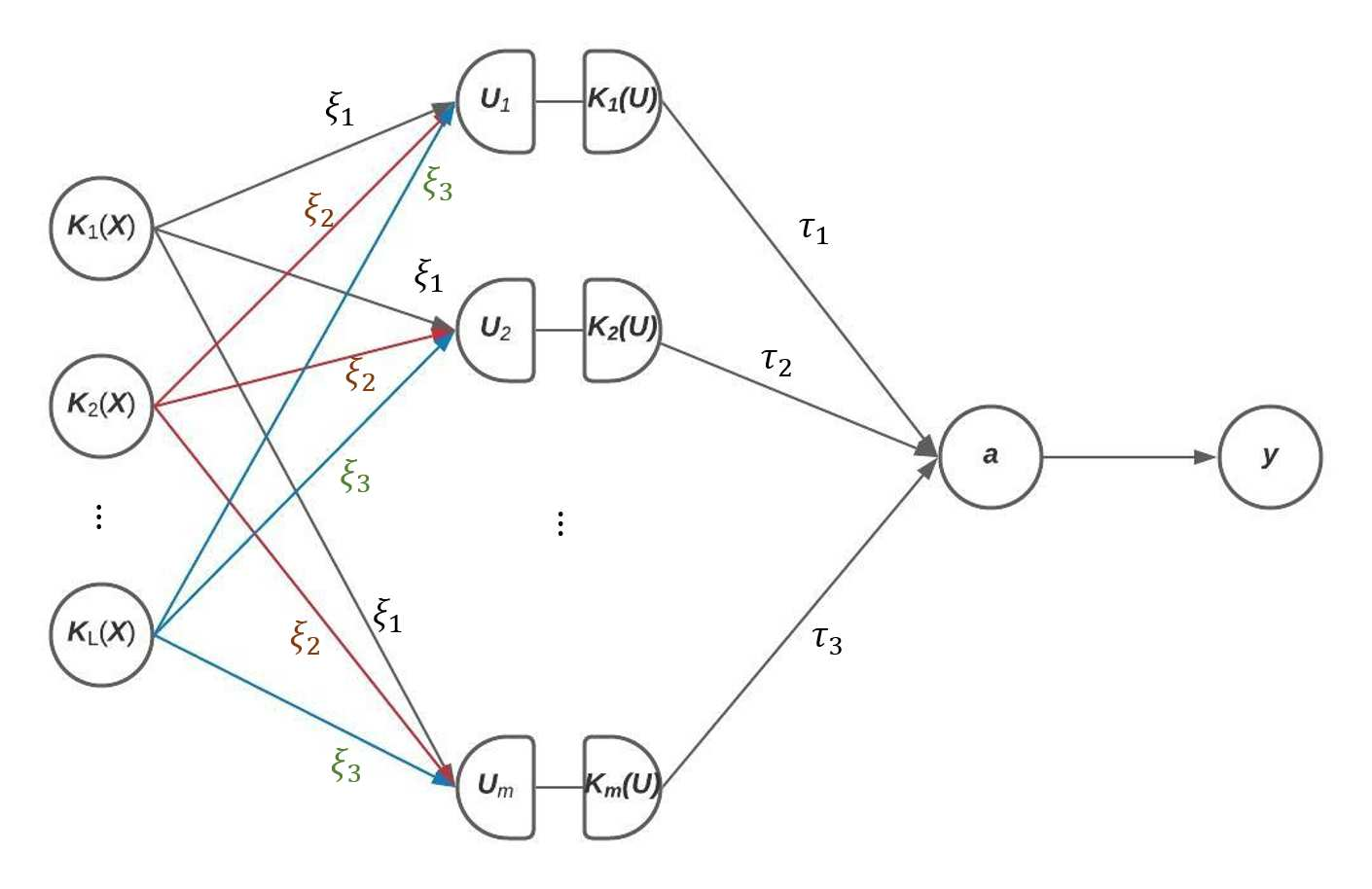
The recent development of artificial intelligence (AI) technology, especially the advance of deep neural network (DNN) technology, has revolutionized many fields. While DNN plays a central role in modern AI technology, it has been rarely used in sequencing data analysis due to challenges brought by high-dimensional sequencing data (e.g., overfitting). Moreover, due to the complexity of neural networks and their unknown limiting distributions, building association tests on neural networks for genetic association analysis remains a great challenge. To address these challenges and fill the important gap of using AI in high-dimensional sequencing data analysis, we introduce a new kernel-based neural network (KNN) test for complex association analysis of sequencing data. The test is built on our previously developed KNN framework, which uses random effects to model the overall effects of high-dimensional genetic data and adopts kernel-based neural network structures to model complex genotype-phenotype relationships. Based on KNN, a Wald-type test is then introduced to evaluate the joint association of high-dimensional genetic data with a disease phenotype of interest, considering non-linear and non-additive effects (e.g., interaction effects). Through simulations, we demonstrated that our proposed method attained higher power compared to the sequence kernel association test (SKAT), especially in the presence of non-linear and interaction effects. Finally, we apply the methods to the whole genome sequencing (WGS) dataset from the Alzheimer's Disease Neuroimaging Initiative (ADNI) study, investigating new genes associated with the hippocampal volume change over time.
翻译:暂无翻译