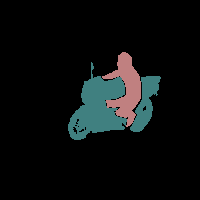
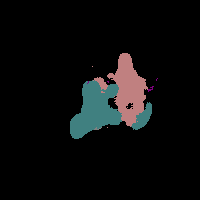
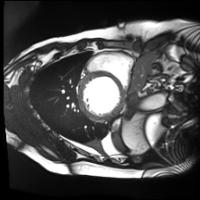
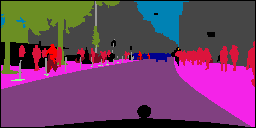
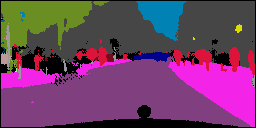
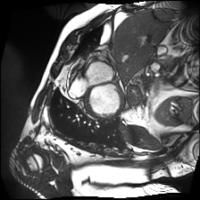
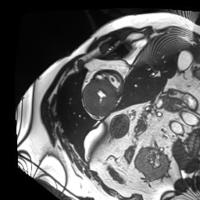
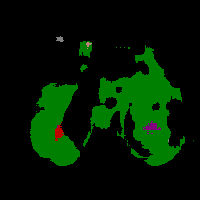
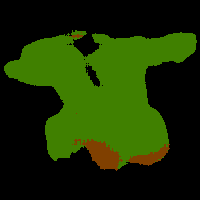
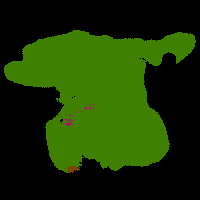
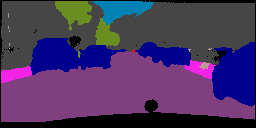
In this work, we study the problem of training deep networks for semantic image segmentation using only a fraction of annotated images, which may significantly reduce human annotation efforts. Particularly, we propose a strategy that exploits the unpaired image style transfer capabilities of CycleGAN in semi-supervised segmentation. Unlike recent works using adversarial learning for semi-supervised segmentation, we enforce cycle consistency to learn a bidirectional mapping between unpaired images and segmentation masks. This adds an unsupervised regularization effect that boosts the segmentation performance when annotated data is limited. Experiments on three different public segmentation benchmarks (PASCAL VOC 2012, Cityscapes and ACDC) demonstrate the effectiveness of the proposed method. The proposed model achieves 2-4% of improvement with respect to the baseline and outperforms recent approaches for this task, particularly in low labeled data regime.
翻译:在这项工作中,我们只使用部分附加说明的图像来培训精密的语义图像分解网络,这可能会大大降低人类的笔记努力。特别是,我们提出一个战略,利用半监督的分解中循环GAN未受保护的图像风格传输能力。与最近使用半监督分解对抗性学习的工程不同,我们强制执行周期一致性,以学习未受监督的图像和分解面遮罩之间的双向绘图。这增加了一种不受监督的正规化效应,在附加说明的数据有限时会提高分解的性能。关于三种不同的公共分解基准(PASCAL VOC 2012, Cityscaccs and ACDC)的实验展示了拟议方法的有效性。拟议模型在基线方面实现了2%至4%的改进,在低标签数据系统中,特别是在低标签数据系统中,比最近这项工作的方法要高出2%至4%。