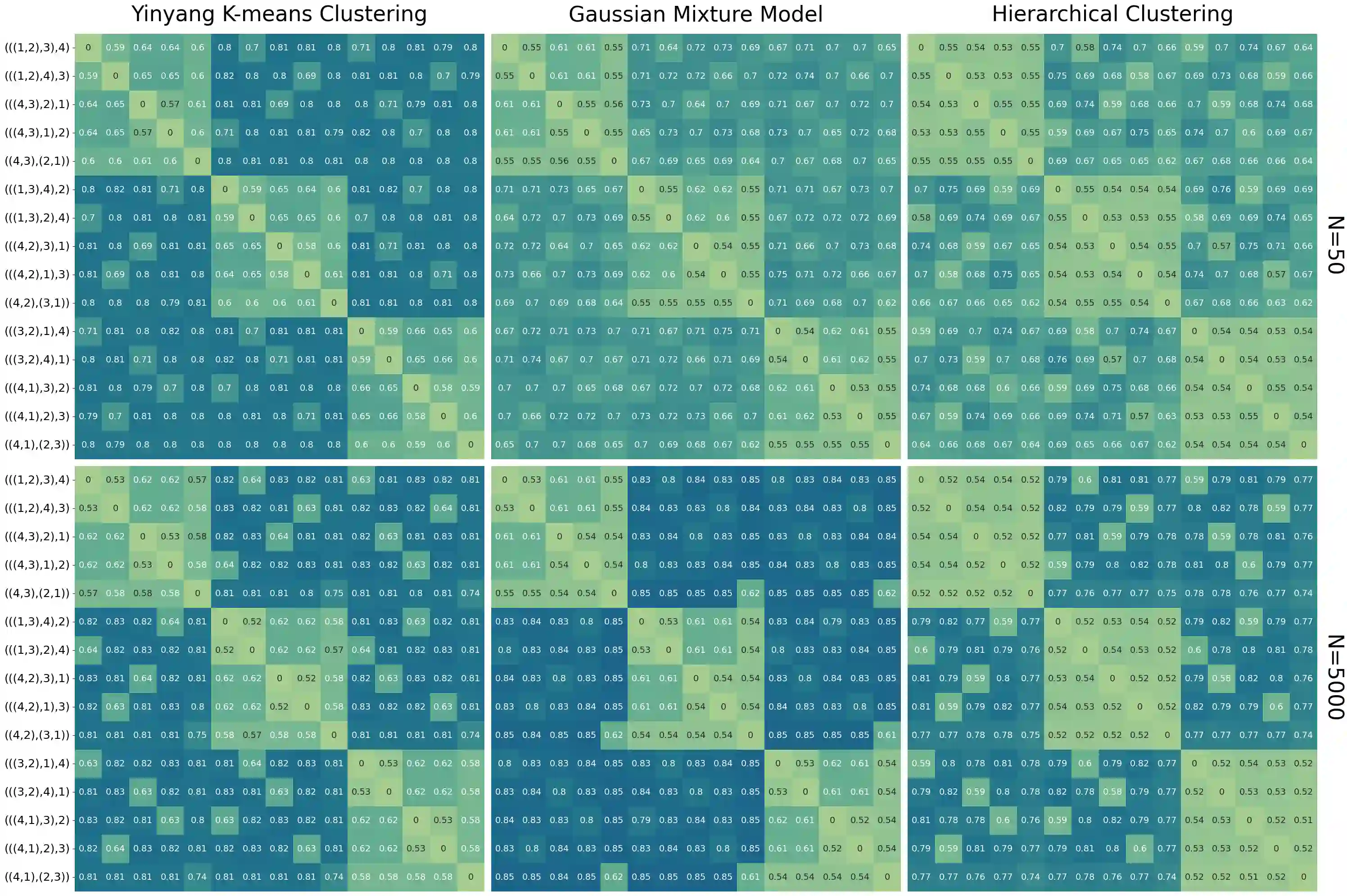
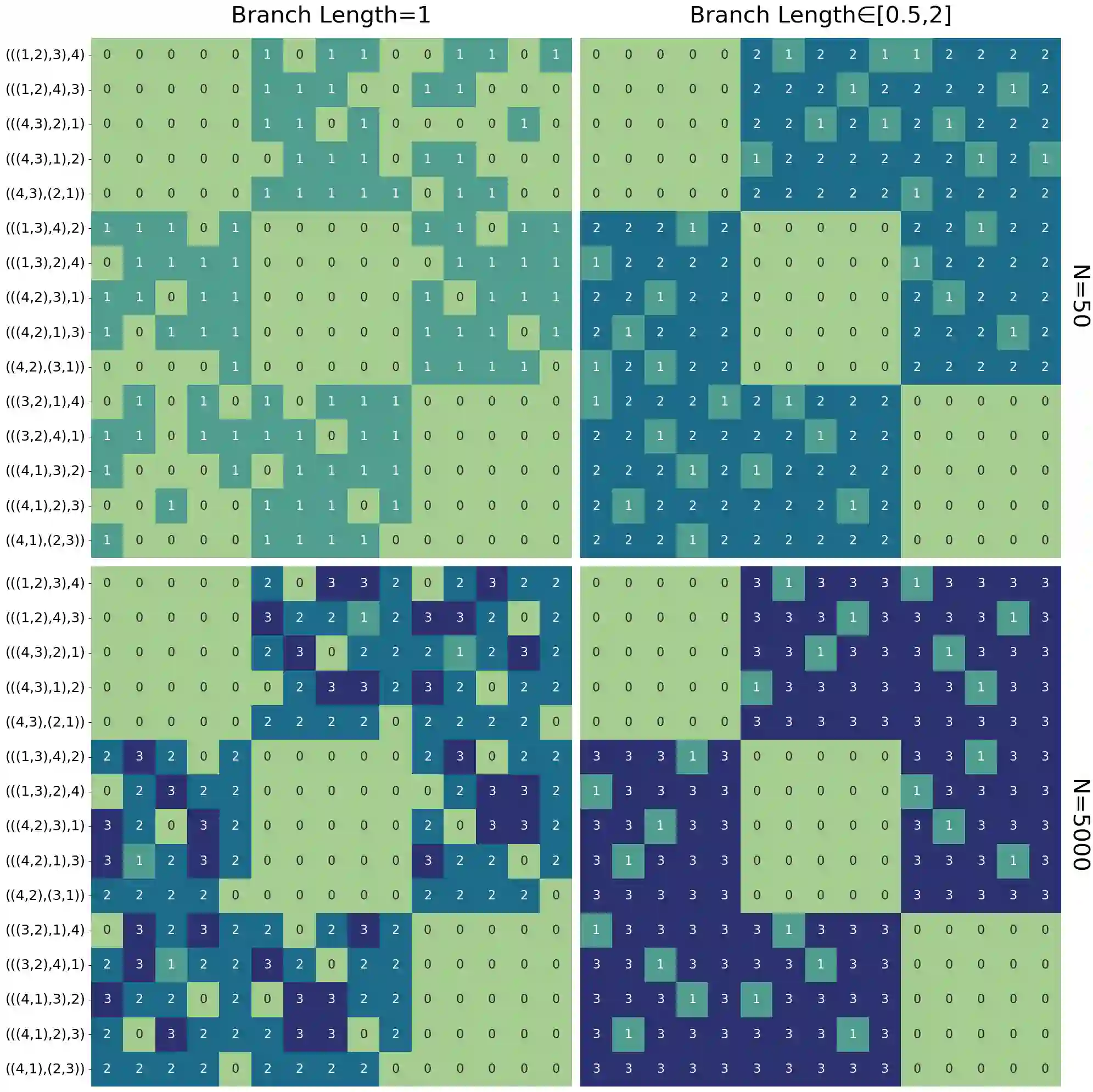
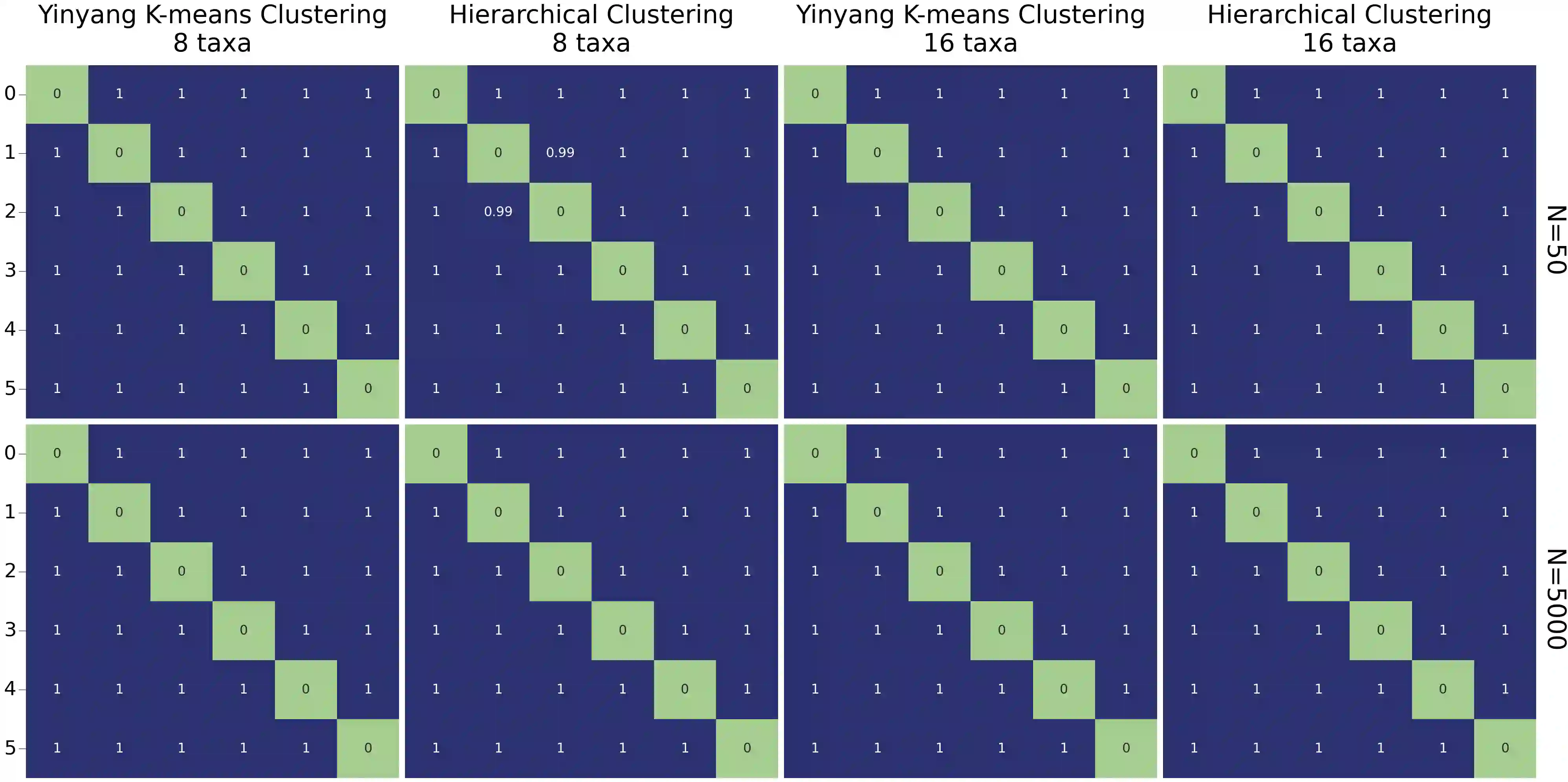
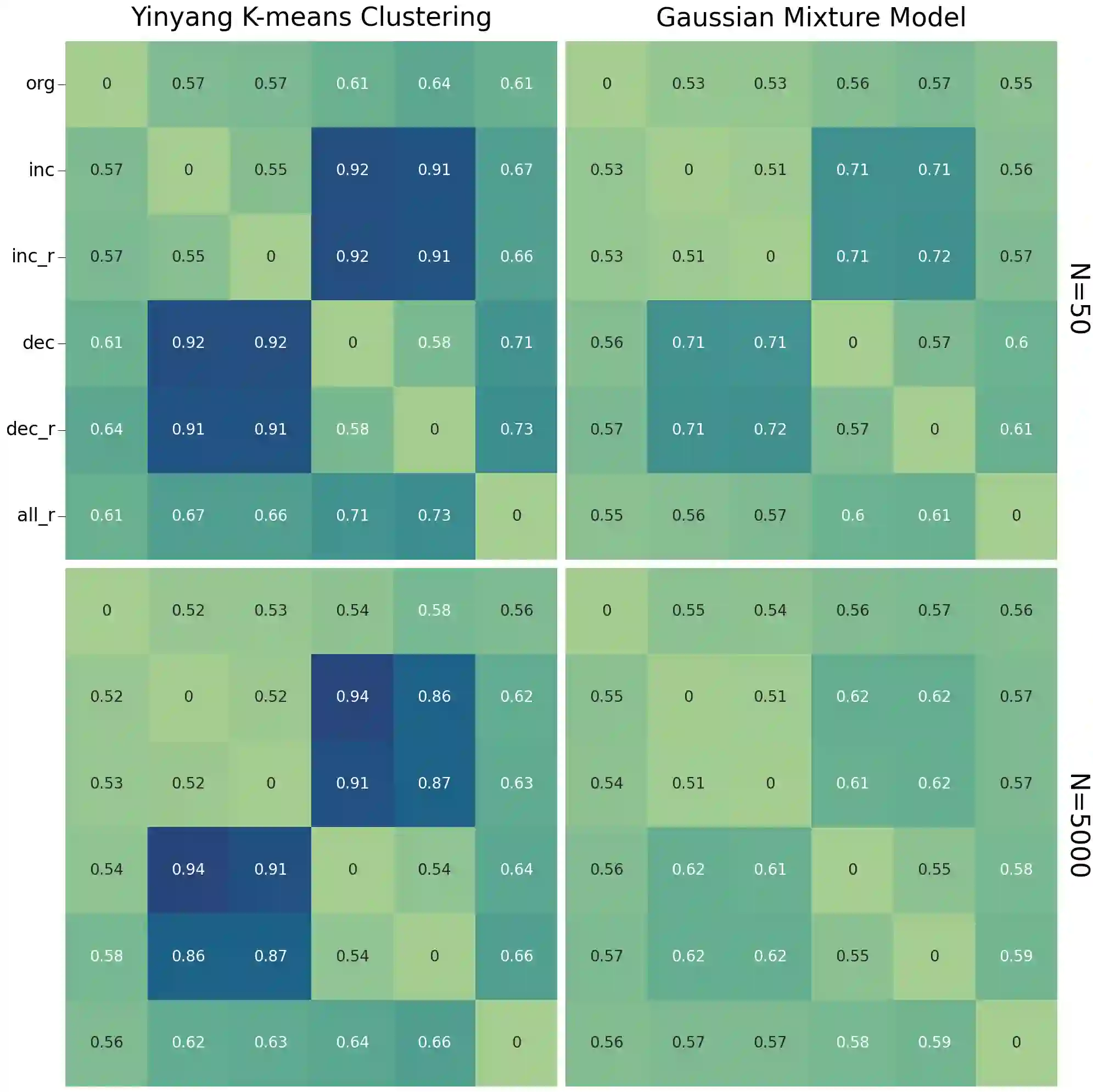
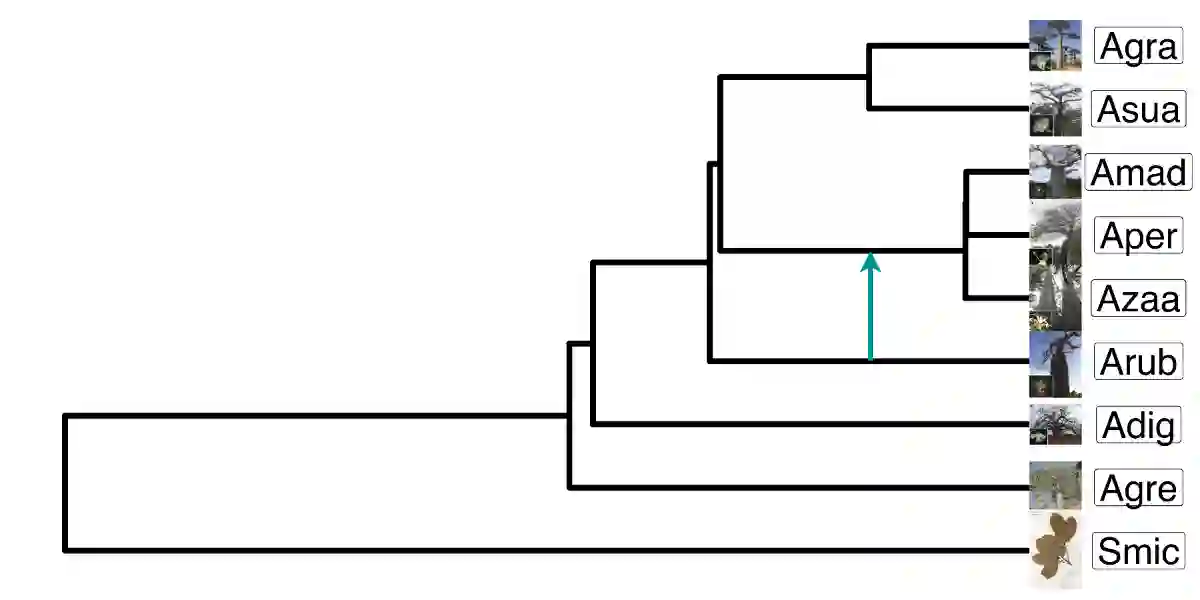
Unsupervised learning has become a staple in classical machine learning, successfully identifying clustering patterns in data across a broad range of domain applications. Surprisingly, despite its accuracy and elegant simplicity, unsupervised learning has not been sufficiently exploited in the realm of phylogenetic tree inference. The main reason for the delay in adoption of unsupervised learning in phylogenetics is the lack of a meaningful, yet simple, way of embedding phylogenetic trees into a vector space. Here, we propose the simple yet powerful split-weight embedding which allows us to fit standard clustering algorithms to the space of phylogenetic trees. We show that our split-weight embedded clustering is able to recover meaningful evolutionary relationships in simulated and real (Adansonia baobabs) data.
翻译:暂无翻译