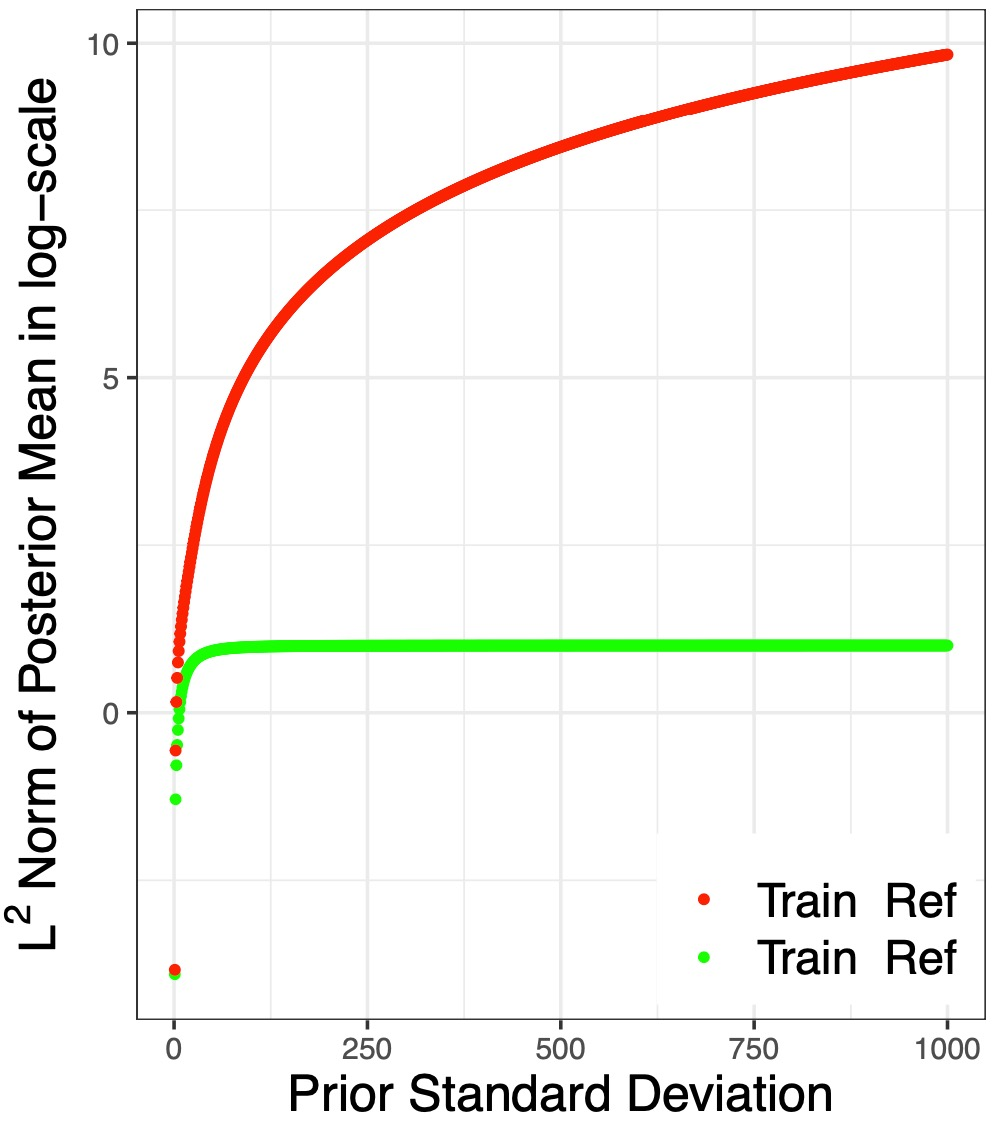
Polygenic Risk Scores (PRS) developed from genome-wide association studies (GWAS) are of increasing interest for various clinical and research applications. Bayesian methods have been particularly popular for building PRS in genome-wide scale because of their natural ability to regularize model and borrow information in high-dimension. In this article, we present new theoretical results, methods, and extensive numerical studies to advance Bayesian methods for PRS applications. We conduct theoretical studies to identify causes of convergence issues of some Bayesian methods when required input GWAS summary-statistics and linkage disequilibrium (LD) (genetic correlation) data are derived from distinct samples. We propose a remedy to the problem by the projection of the summary-statistics data into the column space of the genetic correlation matrix. We further implement a PRS development algorithm under the Bayesian Bridge prior which can allow more flexible specification of effect-size distribution than those allowed under popular alternative methods. Finally, we conduct careful benchmarking studies of alternative Bayesian methods using both simulation studies and real datasets, where we carefully investigate both the effect of prior specification and estimation strategies for LD parameters. These studies show that the proposed algorithm, equipped with the projection approach, the flexible prior specification, and an efficient numerical algorithm leads to the development of the most robust PRS across a wide variety of scenarios.
翻译:暂无翻译