【论文推荐】最新5篇图像分割(Image Segmentation)相关论文—多重假设、超像素分割、自监督、图、生成对抗网络
【导读】专知内容组整理了最近五篇图像分割(Image Segmentation)相关文章,为大家进行介绍,欢迎查看!
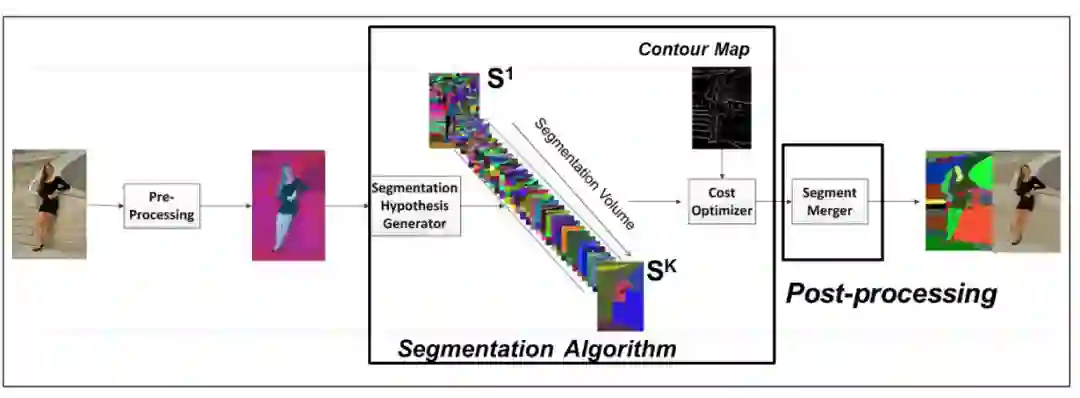
1. Improved Image Segmentation via Cost Minimization of Multiple Hypotheses(通过多重假设最小化损失改进图像分割性能)
作者:Marc Bosch,Christopher M. Gifford,Austin G. Dress,Clare W. Lau,Jeffrey G. Skibo,Gordon A. Christie
摘要:Image segmentation is an important component of many image understanding systems. It aims to group pixels in a spatially and perceptually coherent manner. Typically, these algorithms have a collection of parameters that control the degree of over-segmentation produced. It still remains a challenge to properly select such parameters for human-like perceptual grouping. In this work, we exploit the diversity of segments produced by different choices of parameters. We scan the segmentation parameter space and generate a collection of image segmentation hypotheses (from highly over-segmented to under-segmented). These are fed into a cost minimization framework that produces the final segmentation by selecting segments that: (1) better describe the natural contours of the image, and (2) are more stable and persistent among all the segmentation hypotheses. We compare our algorithm's performance with state-of-the-art algorithms, showing that we can achieve improved results. We also show that our framework is robust to the choice of segmentation kernel that produces the initial set of hypotheses.
期刊:arXiv, 2018年2月1日
网址:
http://www.zhuanzhi.ai/document/dd2ca32cb74a374d653d1e8833bb0177
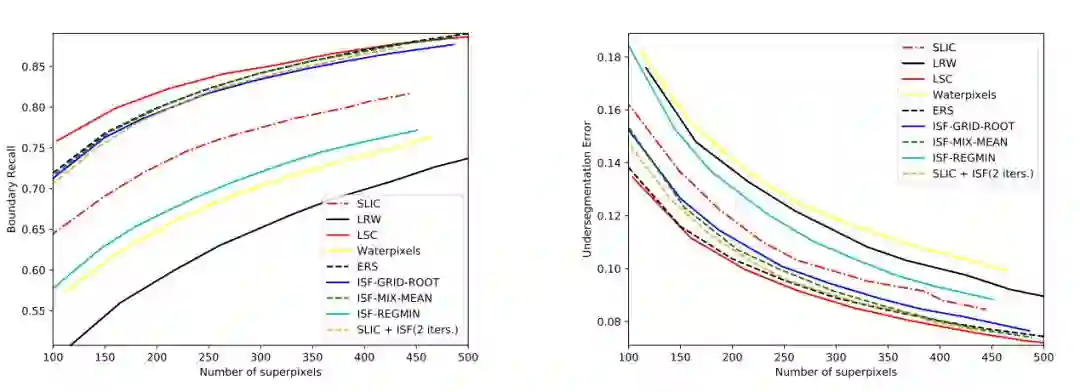
2. An Iterative Spanning Forest Framework for Superpixel Segmentation(一个用于超像素分割的迭代生成森林框架)
作者:John E. Vargas-Muñoz,Ananda S. Chowdhury,Eduardo B. Alexandre,Felipe L. Galvão,Paulo A. Vechiatto Miranda,Alexandre X. Falcão
摘要:Superpixel segmentation has become an important research problem in image processing. In this paper, we propose an Iterative Spanning Forest (ISF) framework, based on sequences of Image Foresting Transforms, where one can choose i) a seed sampling strategy, ii) a connectivity function, iii) an adjacency relation, and iv) a seed pixel recomputation procedure to generate improved sets of connected superpixels (supervoxels in 3D) per iteration. The superpixels in ISF structurally correspond to spanning trees rooted at those seeds. We present five ISF methods to illustrate different choices of its components. These methods are compared with approaches from the state-of-the-art in effectiveness and efficiency. The experiments involve 2D and 3D datasets with distinct characteristics, and a high level application, named sky image segmentation. The theoretical properties of ISF are demonstrated in the supplementary material and the results show that some of its methods are competitive with or superior to the best baselines in effectiveness and efficiency.
期刊:arXiv, 2018年1月30日
网址:
http://www.zhuanzhi.ai/document/84d288b969678cd6a42661ad0ffb0a03
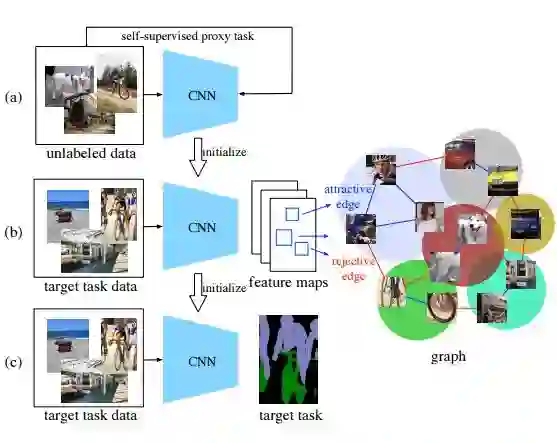
3. Mix-and-Match Tuning for Self-Supervised Semantic Segmentation(自监督语义分割的混合匹配优化)
作者:Xiaohang Zhan,Ziwei Liu,Ping Luo,Xiaoou Tang,Chen Change Loy
摘要:Deep convolutional networks for semantic image segmentation typically require large-scale labeled data, e.g. ImageNet and MS COCO, for network pre-training. To reduce annotation efforts, self-supervised semantic segmentation is recently proposed to pre-train a network without any human-provided labels. The key of this new form of learning is to design a proxy task (e.g. image colorization), from which a discriminative loss can be formulated on unlabeled data. Many proxy tasks, however, lack the critical supervision signals that could induce discriminative representation for the target image segmentation task. Thus self-supervision's performance is still far from that of supervised pre-training. In this study, we overcome this limitation by incorporating a "mix-and-match" (M&M) tuning stage in the self-supervision pipeline. The proposed approach is readily pluggable to many self-supervision methods and does not use more annotated samples than the original process. Yet, it is capable of boosting the performance of target image segmentation task to surpass fully-supervised pre-trained counterpart. The improvement is made possible by better harnessing the limited pixel-wise annotations in the target dataset. Specifically, we first introduce the "mix" stage, which sparsely samples and mixes patches from the target set to reflect rich and diverse local patch statistics of target images. A "match" stage then forms a class-wise connected graph, which can be used to derive a strong triplet-based discriminative loss for fine-tuning the network. Our paradigm follows the standard practice in existing self-supervised studies and no extra data or label is required. With the proposed M&M approach, for the first time, a self-supervision method can achieve comparable or even better performance compared to its ImageNet pre-trained counterpart on both PASCAL VOC2012 dataset and CityScapes dataset.
期刊:arXiv, 2018年1月30日
网址:
http://www.zhuanzhi.ai/document/42991861993f309f7e7bc7fcad635556
4. Deep LOGISMOS: Deep Learning Graph-based 3D Segmentation of Pancreatic Tumors on CT scans(Deep LOGISMOS: CT扫描中胰腺肿瘤的深度学习图3D分割方法)
作者:Zhihui Guo,Ling Zhang,Le Lu,Mohammadhadi Bagheri,Ronald M. Summers,Milan Sonka,Jianhua Yao
摘要:This paper reports Deep LOGISMOS approach to 3D tumor segmentation by incorporating boundary information derived from deep contextual learning to LOGISMOS - layered optimal graph image segmentation of multiple objects and surfaces. Accurate and reliable tumor segmentation is essential to tumor growth analysis and treatment selection. A fully convolutional network (FCN), UNet, is first trained using three adjacent 2D patches centered at the tumor, providing contextual UNet segmentation and probability map for each 2D patch. The UNet segmentation is then refined by Gaussian Mixture Model (GMM) and morphological operations. The refined UNet segmentation is used to provide the initial shape boundary to build a segmentation graph. The cost for each node of the graph is determined by the UNet probability maps. Finally, a max-flow algorithm is employed to find the globally optimal solution thus obtaining the final segmentation. For evaluation, we applied the method to pancreatic tumor segmentation on a dataset of 51 CT scans, among which 30 scans were used for training and 21 for testing. With Deep LOGISMOS, DICE Similarity Coefficient (DSC) and Relative Volume Difference (RVD) reached 83.2+-7.8% and 18.6+-17.4% respectively, both are significantly improved (p<0.05) compared with contextual UNet and/or LOGISMOS alone.
期刊:arXiv, 2018年1月26日
网址:
http://www.zhuanzhi.ai/document/deab2fa98863fa8b5011c2df13f6ec31
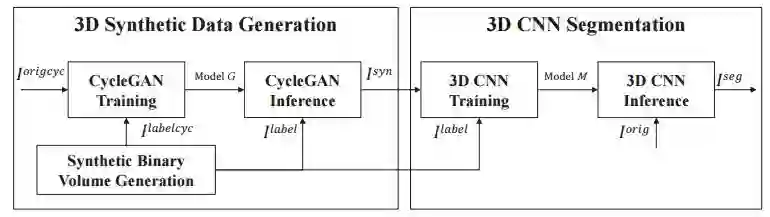
5. Fluorescence Microscopy Image Segmentation Using Convolutional Neural Network With Generative Adversarial Networks(基于卷积神经网络与生成对抗网络的荧光显微图像分割)
作者:Chichen Fu,Soonam Lee,David Joon Ho,Shuo Han,Paul Salama,Kenneth W. Dunn,Edward J. Delp
摘要:Recent advance in fluorescence microscopy enables acquisition of 3D image volumes with better quality and deeper penetration into tissue. Segmentation is a required step to characterize and analyze biological structures in the images. 3D segmentation using deep learning has achieved promising results in microscopy images. One issue is that deep learning techniques require a large set of groundtruth data which is impractical to annotate manually for microscopy volumes. This paper describes a 3D nuclei segmentation method using 3D convolutional neural networks. A set of synthetic volumes and the corresponding groundtruth volumes are generated automatically using a generative adversarial network. Segmentation results demonstrate that our proposed method is capable of segmenting nuclei successfully in 3D for various data sets.
期刊:arXiv, 2018年1月23日
网址:
http://www.zhuanzhi.ai/document/9fdb6962bd4223b7ad1e6d818a0a304c
-END-
专 · 知
人工智能领域主题知识资料查看获取:【专知荟萃】人工智能领域26个主题知识资料全集(入门/进阶/论文/综述/视频/专家等)
同时欢迎各位用户进行专知投稿,详情请点击:
【诚邀】专知诚挚邀请各位专业者加入AI创作者计划!了解使用专知!
请PC登录www.zhuanzhi.ai或者点击阅读原文,注册登录专知,获取更多AI知识资料!
请扫一扫如下二维码关注我们的公众号,获取人工智能的专业知识!
请加专知小助手微信(Rancho_Fang),加入专知主题人工智能群交流!
点击“阅读原文”,使用专知!