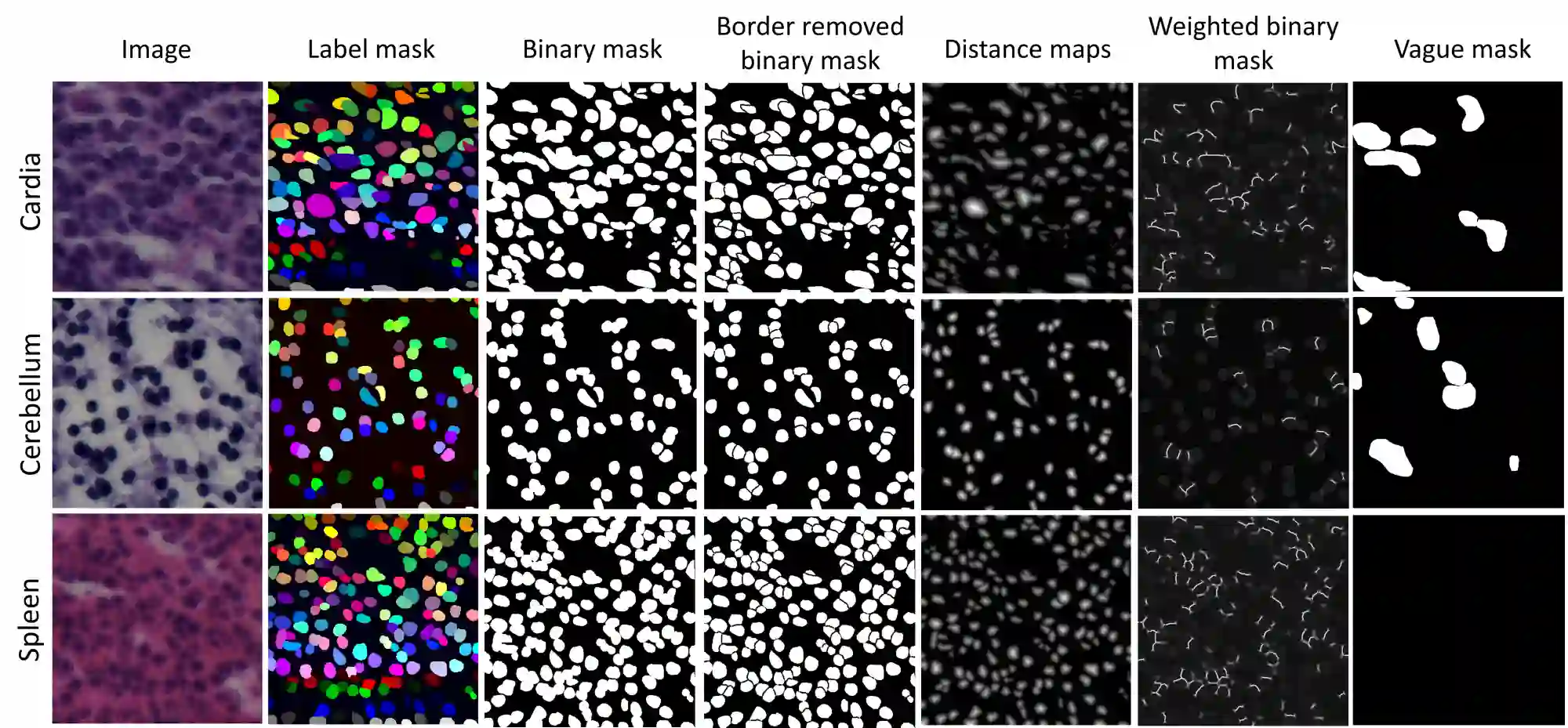
In computational pathology, automatic nuclei instance segmentation plays an essential role in whole slide image analysis. While many computerized approaches have been proposed for this task, supervised deep learning (DL) methods have shown superior segmentation performances compared to classical machine learning and image processing techniques. However, these models need fully annotated datasets for training which is challenging to acquire, especially in the medical domain. In this work, we release one of the biggest fully manually annotated datasets of nuclei in Hematoxylin and Eosin (H&E)-stained histological images, called NuInsSeg. This dataset contains 665 image patches with more than 30,000 manually segmented nuclei from 31 human and mouse organs. Moreover, for the first time, we provide additional ambiguous area masks for the entire dataset. These vague areas represent the parts of the images where precise and deterministic manual annotations are impossible, even for human experts. The dataset and detailed step-by-step instructions to generate related segmentation masks are publicly available at https://www.kaggle.com/datasets/ipateam/nuinsseg and https://github.com/masih4/NuInsSeg, respectively.
翻译:暂无翻译